Design de novo antibody-antigen interactions with RFantibody
Designing an antibody from scratch to bind a specific site on a target protein has traditionally required immunization or large-scale library screening. But now, with the rise of deep learning tools, it’s possible to computationally generate antibodies tailored to precise epitopes—from the ground up.
One of the most exciting advancements in this space is RFantibody, a deep learning-based framework from the Baker lab at University of Washington for enhanced design and prediction of antibodies to a target. We are pleased to announce that the full RFantibody pipeline is available now at Levitate Bio!
What is RFantibody: A step-by-step guide
RFantibody is an end-to-end antibody design pipeline that leverages deep learning and molecular modeling to create novel antibody structures with enhanced binding affinity and specificity. It contains 3 tools for generating de novo antibodies bound to a protein target. Additionally, Levitate has tools to help identify and optimize hits that complement these protocols.
-
RFdiffusion fine-tuned on antibody complex structures for generating complete antibodies in the bound state.
-
ProteinMPNN to design sequences that fold into your RFdiffusion antibodies and retain the bound mode.
-
Fine-tuned RoseTTAFold2 for antibody-antigen prediction for validating which ProteinMPNN sequences fold into the bound mode as expected.
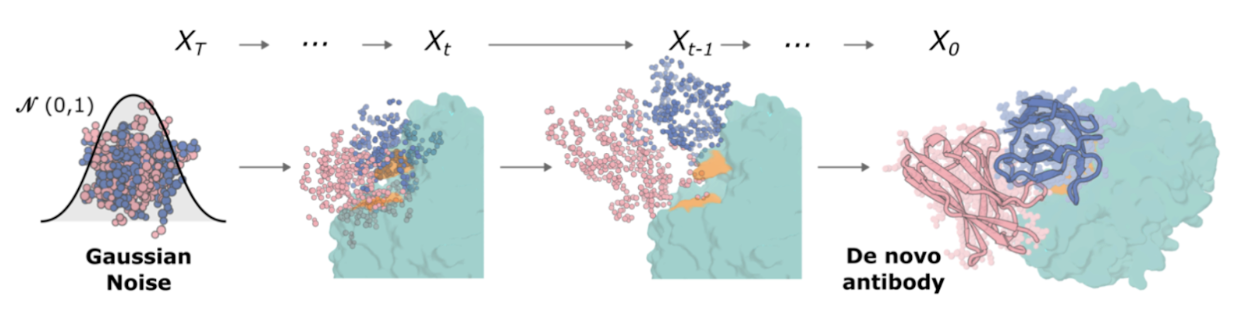
Step one: Fine-tuned RFdiffusion
RFdiffusion is a deep-learning framework for designing proteins. It is based on a diffusion model, similar to those used in generative AI for images (like Stable Diffusion), but instead of generating images, it generates novel protein structures.
Background
RFdiffusion uses a denoising diffusion probabilistic model (DDPM) to iteratively refine and generate protein structures. RFantibody RFdiffusion was trained on a wide range of antibody-antigen complexes. It learned how to understand these interactions by gradually corrupting existing protein structures with random noise. Then it optimized its algorithm so it could reverse this process, reconstructing the original protein structures step by step. Therefore, if you give it a new antigen, it can create a new antibody in a bound mode.
De novo antibody structure generation
1) Protein Generation via Diffusion Starting from a randomly generated or partially specified antibody backbone, the model iteratively removes noise while shaping the structure into a physically plausible protein.
2) Conditioning on Constraints The model can be guided to generate proteins with specific shapes, binding sites, or functions by conditioning the diffusion process on desired structural features.
For example, our general purpose tool for RFdiffusion can design proteins that bind to a specific target or form symmetric oligomers and RFdiffusion all atom can design protein with ligands and nucleic acids present.
3) Sequence Design and Structure Validation After generation, the designed proteins are further refined using ProteinMPNN sequence design and structure prediction with AlphaFold or Boltz before being experimentally validated to ensure they fold correctly and function as intended.
The RFantibody model for RFdiffusion allows users to customize the antigen and antibody information provided for a highly precise design task, or more unspecified interactions.
- The epitope to target (shown below as surface).
- Which CDR loops to redesign.
- An antibody framework (VHH or scFv).
- The number of antibody designs to generate.
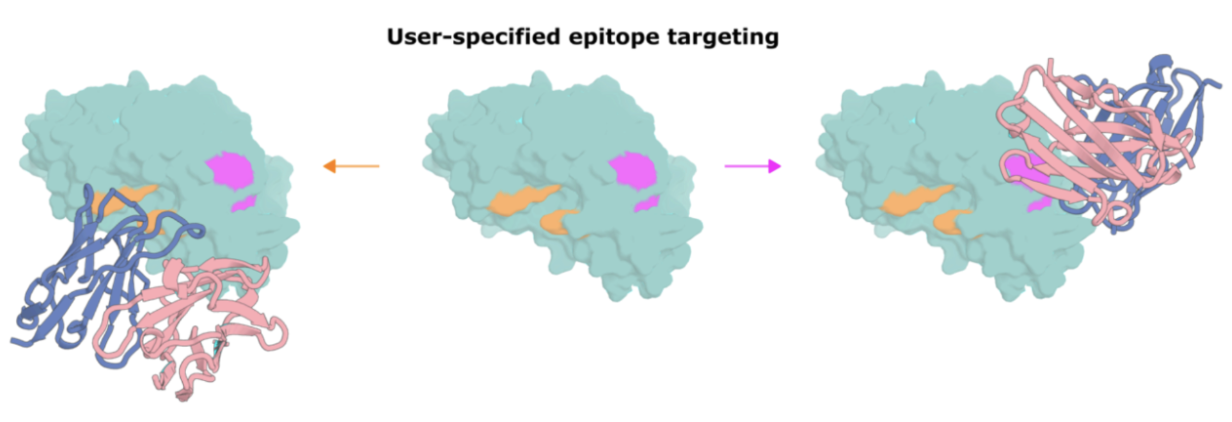
Step two: ProteinMPNN
ProteinMPNN is a deep learning-based model that takes a fixed protein backbone (3D structure) as input and predicts the most likely amino acid sequence that will fold into that structure.
Background
The model receives the 3D coordinates of a protein backbone (without knowing the amino acid sequence). The backbone consists of only the spatial arrangement of alpha-carbon (Cα) atoms and the main chain of the protein.
Based on the structural context provided by the backbone, ProteinMPNN uses graph neural network (GNN) processing to predict the optimal amino acid for each position. The sequence is optimized to be thermodynamically stable and functionally relevant.
Antibody sequence design
ProteinMPNN for RFantibody has no additional fine-tuning. However, it limits sequence design to the CDRs for efficient automation. Simply pass in structures from fine-tuned RFDiffusion to RFantibody ProteinMPNN and it will give you sequences for the predicted CDR sequences in return. If you want to design sequence on more than just the CDRs, you can use our general purpose ProteinMPNN tool. These sequences can then be validated structurally with fine-tuned RosettaFold2.
Step three: Fine-tuned RoseTTAFold2
RoseTTAFold2 is an advanced deep-learning model for protein structure prediction and design. It is an evolution of the original RoseTTAFold, incorporating ideas from AlphaFold2 but optimized for speed, accuracy, and modularity in protein modeling.
Background
RoseTTAFold2 generates a high-confidence atomic-resolution 3D model of a protein, even modeling protein-protein interactions and multi-chain assemblies, making it an ideal base model for antibody complexes.
RFantibody RoseTTAFold2 is fine-tuned on antibody complexes, making it capable of recapitulating experimentally determined antibody framework structures and distinguishing between real and decoy antibody-antigen interactions.
Designed antibody validation in silico
To use RFantibody RoseTTAFold2, simply pass in designs from RFantibody ProteinMPNN as input and this tool will give you bound complex predictions of your novel antibody and antigen. The lower the RMSD of the RoseTTAFold2 prediction to the RFdiffusion model is, the better the structure and sequence design.
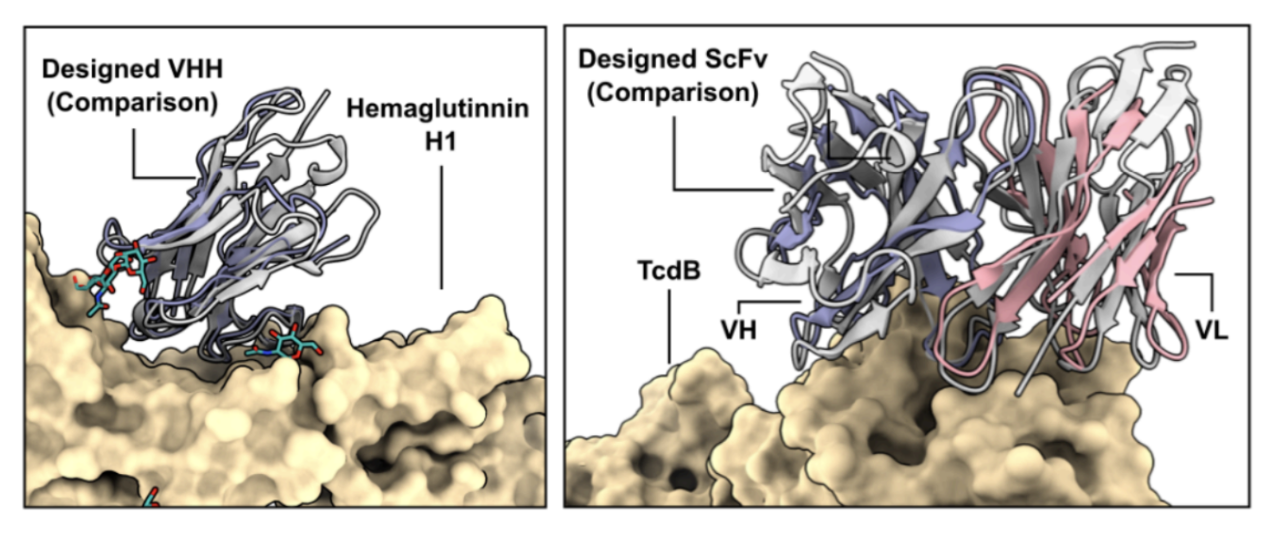
A designed VHH to hemaglutinin and a designed scFv to TcdB were characterized with cryo-EM and the RFantibody design (grey) was overlaid with the experimental structure (purple and pink). The RMSD of the experimental and designed structures was 1.45 Å and 0.9 Å, respectively. The close alignment of the structures shows how accurately the RFantibody pipeline represents the antibody framework and CDRs.
How many antibodies should you test?
Designing de novo antibody-based binders more challenging than designing non-antibody binders, due to the complex loop architecture and strict framework constraints. To improve your chances of success, we recommend the following strategy:
- Generate a large number of antibody structures using Antibody RFdiffusion.
- Sample diverse backbones, then use ProteinMPNN to design multiple sequences for each backbone.
- Filter sequences using the fine-tuned RoseTTAFold2 model, keeping only those predicted to bind in the expected orientation.
- Apply additional filters from Levitate, such as:
- Solubility scoring to predict developability.
- Interface Analyzer to score the antibody-antigen interface for affinity and specificity.
This workflow helps you prioritize designs with the highest likelihood of folding correctly, being soluble, and binding the target epitope with strong affinity.
Once you’ve filtered your top candidates, the number to test in the wet lab depends on your screening approach:
- Low-throughput (e.g., SPR, ELISA): Start with 100–200 designs per epitope
- High-throughput (e.g., yeast display): Test 1,000–10,000 designs for broad coverage
Top binders can then be affinity matured or further optimized using Levitate’s Rosetta-based design tools.
Takeaways
RFantibody is a powerful pipeline for de novo antibody design, allowing researchers to create a novel antibody-antigen interaction (RFdiffusion) and determine a sequence that delivers the intended fold (ProteinMPNN). Finally, antibody-antigen complex validation (RoseTTAFold2) and Levitate scoring tools indicate developability and high affinity binding, allowing you to choose only the best designs for experimental testing.
RFantibody is now part of our tool set at Levitate Bio and we would be thrilled to help you introduce it into your workflows. If you’d like to try designing de novo antibodies with RFantibody, don’t hesitate to reach out to us!
References
Nathaniel Bennett, Joseph Watson, Robert Ragotte, David Baker, et al. Atomically accurate de novo design of antibodies with RFdiffusion. https://doi.org/10.1101/2024.03.14.585103