Engineering a TCR Mimic that binds a target cancer-associated peptide:MHC complex with strong peptide specificity
A Real-World Example of Antibody Design
A recent BioRxiv preprint by Oh et al. highlights the potential of peptide–HLA (pHLA) complexes that present disease-specific peptides as powerful therapeutic targets. TCR-mimetic antibodies that recognize an antigen only in its pHLA context can precisely eliminate diseased cells, but the strategy is complicated by extensive HLA polymorphism, which requires high peptide specificity with minimal HLA engagement. As an example, consider the Fab–pHLA complex bound to the cancer-associated peptide RMFPNAPYL.
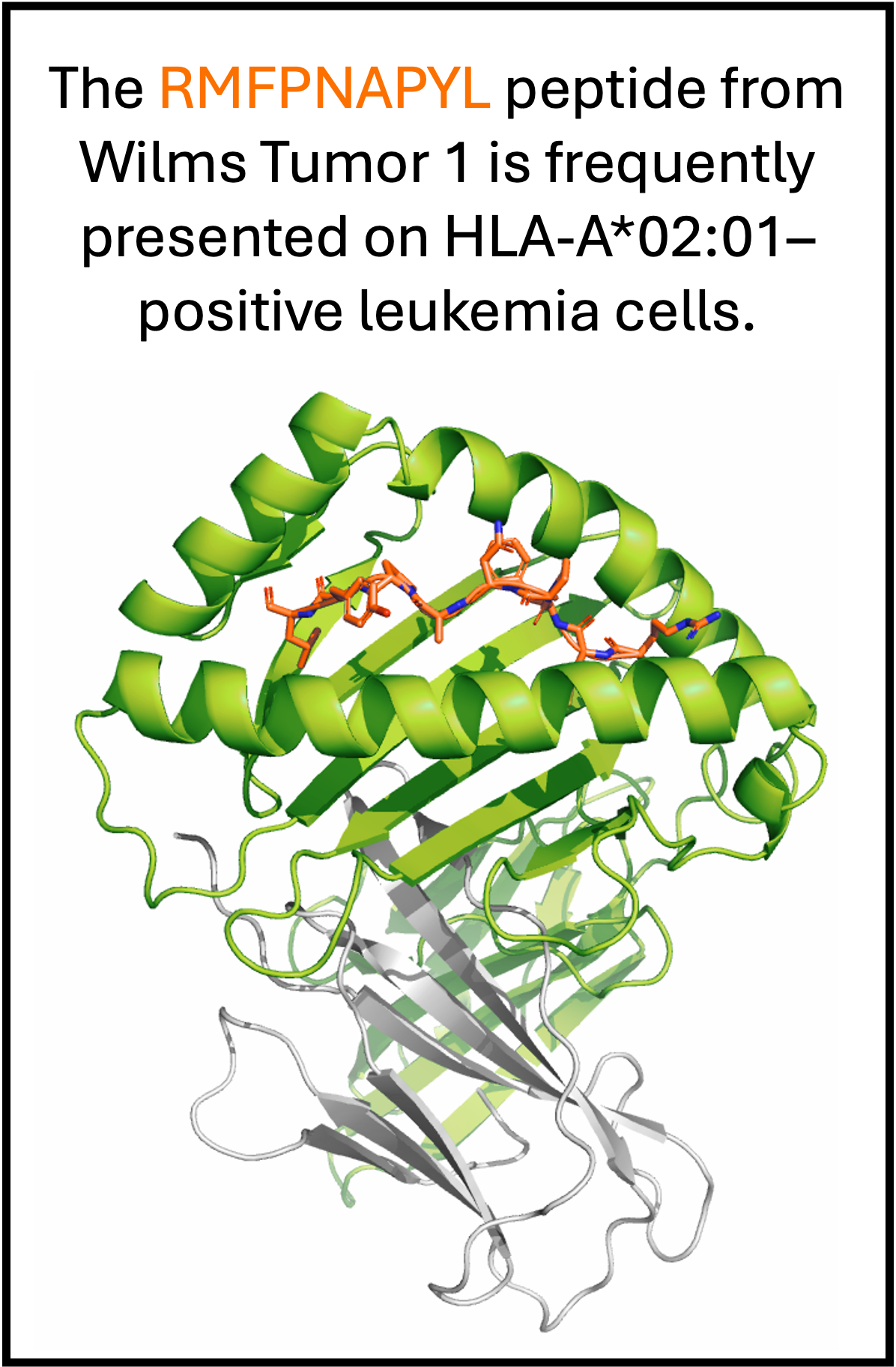
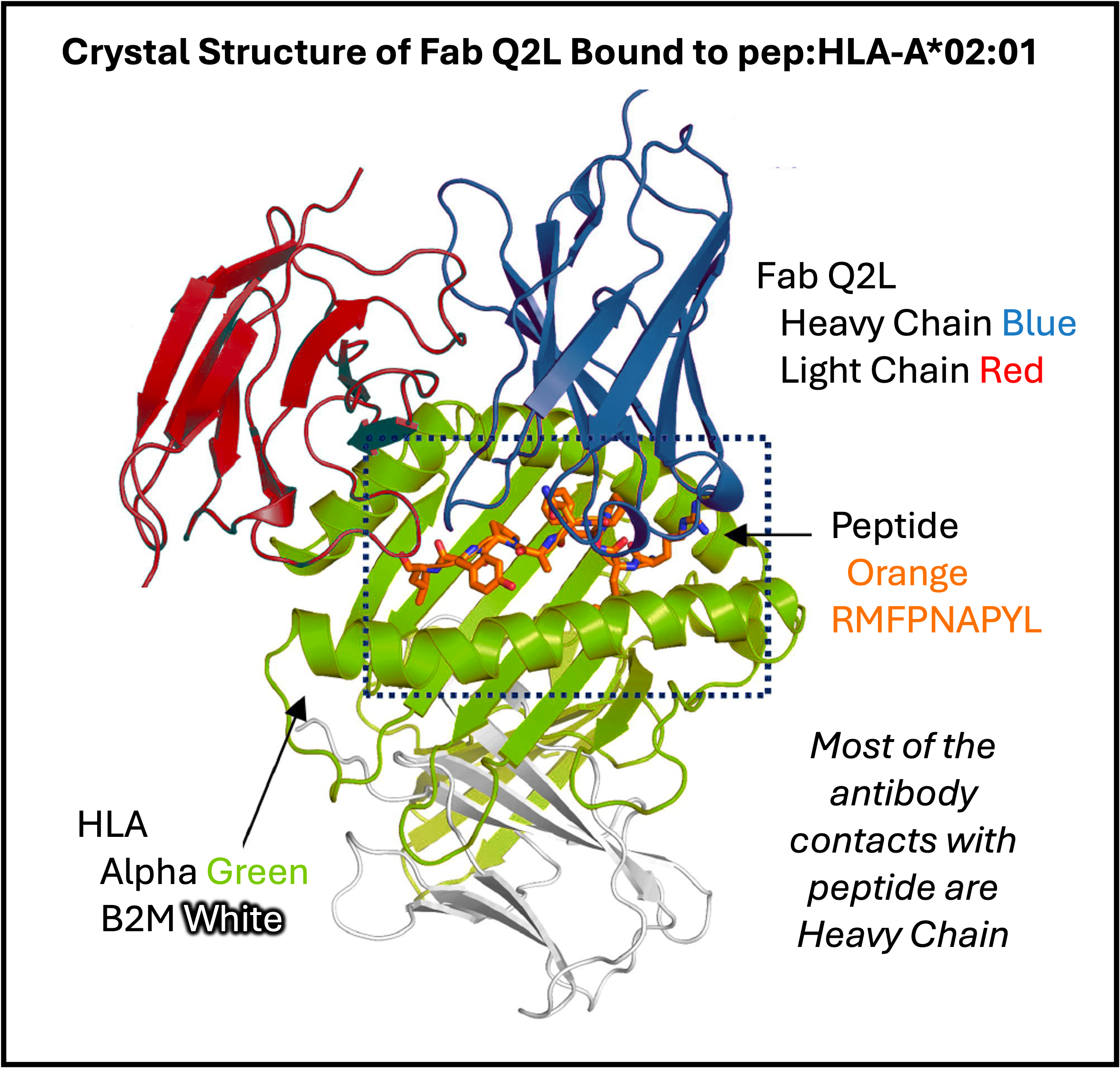
The crystal structure of Fab Q2L bound to RMFPNAPYL/HLA-A*02:01 showed that many of the Fab contacts are in the heavy chain CDRs, inspiring the authors to engineer a single-domain VH that strengthens peptide binding while minimizing HLA contacts, yielding a peptide-specific heavy chain.
Since the Q2L Fab’s VH is not naturally stable without the light chain, they used a known soluble human VH scaffold from Trastuzumab (Tra_VH, engineered with 5 mutations for solubility) as the starting framework. They grafted full H3 and transplanted some of H1 and H2 from Q2L and created sequence modifications to remove clashes, improve affinity to the peptide, and to generate favorable features in the chimera for developability using Rosetta to generate the novel design and Chai to validate the bound structure.
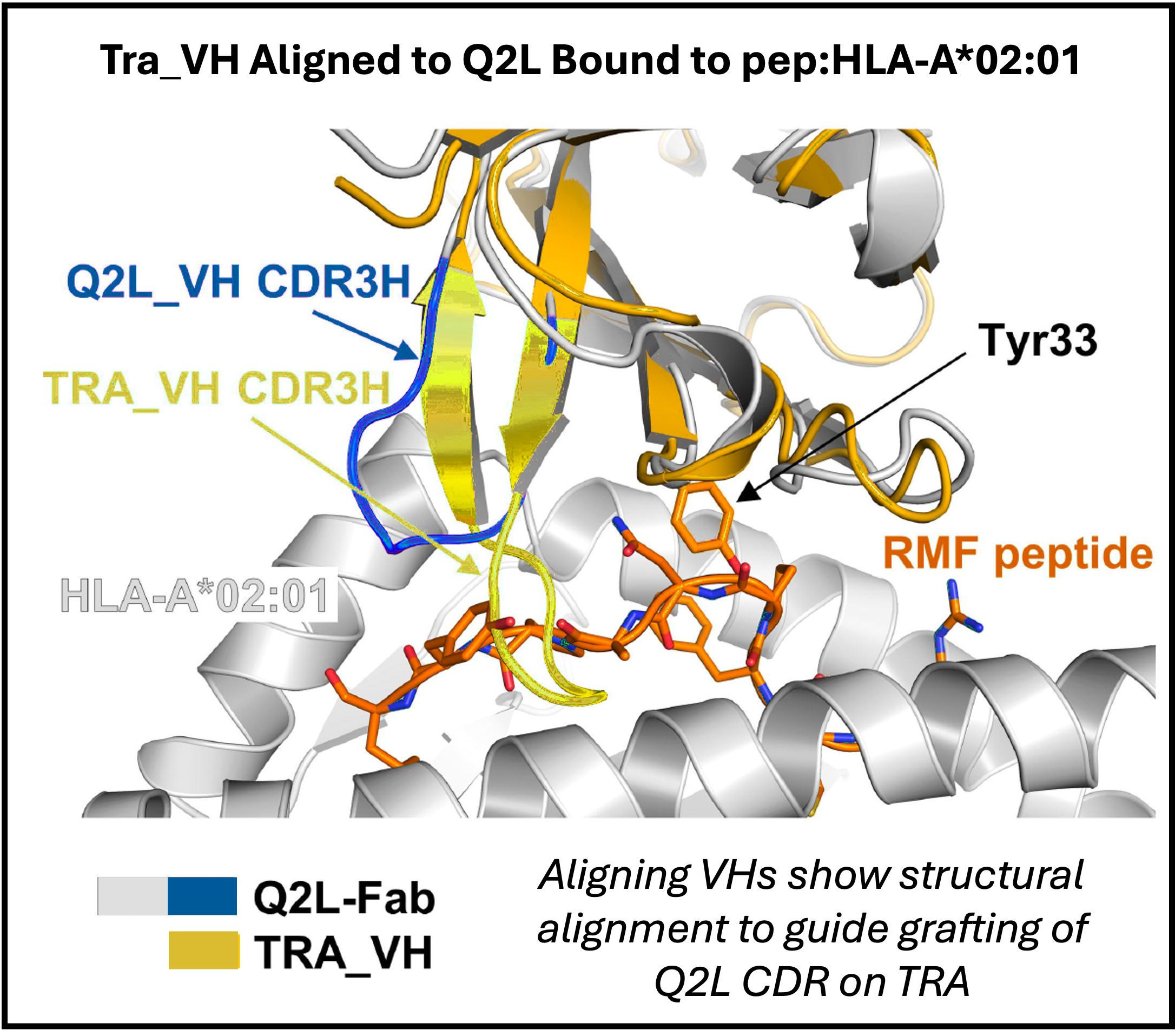
The best design had 7 point mutations and a 4-residue deletion relative to Tra_VH. The model predicted retention of key peptide contacts and minimal HLA engagement. Wet lab validated showed strong, specific binding (Flow Cytometry validated binding on T2 cells with RMF, incorporation with Car-T showed dose-dependent IL-2 production, and BLI measured KD 0.37 nM for bivalent form ~200× affinity improvement).
In the following section, we will walk through how you can accomplish a similar antibody design pipeline using automated APIs in the Levitate Engine platform. These tools are hand-selected for their accuracy, speed, and reliability. To see a live demo of this pipeline and any others, contact info@levitate.bio.
Walkthrough of Antibody Design with Levitate Tools
1) Structure prediction of the HLA with the peptide bound to the antibody (if you don’t have a crystal structure).
-
Colab Search - create a multi-sequence alignment (MSA) for both the HLA and antibody.
-
Boltz - run structure prediction with these MSAs and their respective input sequences
-
Tip: Boltz can be run with defined binding site residues to increase the chance of obtaining an accurate complex prediction.
-
If predicted accuracy is low, consider hiring us to run TCRModel2 which has a mode for antibody-peptide-HLA.
2) Structure optimization - Using structure predictions with high confidence or experimentally obtained structures, optimize the structure by performing conformational sampling and scoring.
-
Rosetta Relax** explores the conformational space by allowing backbones and side chains to shift and rotate to find more favorable states.
-
Tip: Generate multiple conformations (eg. repeats=20) to distinguish dynamics regions vs where the structure is already in the most favorable conformation.
3) Create chimeras - key sequences from CDRs of the input antibody replace VH sequences to create chimeric sequences.
- Design will take custom sequence sampling parameters as input to design chimeric sequences.
4) Structure prediction of chimeric antibodies can be done in multiple ways with Levitate tools.
-
Immunebuilder can do rapid, high throughput structure prediction if you have generated hundreds or thousands of chimeric sequences.
-
Tip: You may filter out sequences in which the CDRs don’t have high confidence. Top hits can also be aligned to the heavy chain in the antibody-peptide-HLA structure - we can write a script for you that will run Rosetta or BioPython and can score CDR alignments and output bound modes for further analysis and/or design.
-
Boltz can take the antibody-peptide-HLA structure with your chimeric sequence to generate a bound mode.
5) Analysis of sequences and structures can be done to rank or to identify regions to run design
-
BioPhi scores sequences by human-likeness, one way of analyzing potential immunogenicity.
-
Epitope Scan will predict binding likelihood of all sequence regions to most common class II MHCs, which indicates how likely the sequence is to be displayed to the immune system.
-
Solubility scoring is a measure of propensity for aggregation to rank structures. It also identifies positions in the protein that are likely to contribute to potential aggregation.
-
PTM Prediction does sequence and structure-based prediction of positions likely to have 9 PTMs that are associated with degradation when scaling up in manufacturing of protein therapeutics.
-
Rosetta Interface Analysis scores the antibody interface with the peptide and/or the HLA for a broad range of features such as interface energy, number of interface of contact, hydrogen bonds, etc.
-
DeepViscosity provides predictions antibody sequence viscosity.
6) Antibody Design - We have Rosetta and AI approaches to find more favorable sequences.
-
ThermoMPNN - Single point mutation scan of every position of the chimeric structures can be done to identify favorable single mutations.
-
Rosetta’s Relax Design - Select regions for mutation sampling and generate libraries of sequence improvements that can often identify combinations of mutations that improve stability, solubility, affinity/selectivity, or to remove liabilities. You can select custom sequence bias of exclusions for individual positions and you can guide general features like max # of mutations, # of charged mutations, etc. Best sequences for bound-state designs should be scored in unbound state.
-
BioPhi - this tool can take chimeric sequences and the structure or the new antibody in bound or unbound modes and you can set your objectives, which can be to have good humanness score and good developability. You can fully lock positions or allow modest amount of changes and you can bias sequence type. Like Relax Design, BioPhi can balance many factors and score bound and unbound states to help select final hits.
In summary, Levitate tools can replicate a TCR-mimic antibody design workflow by guiding users through structure prediction, optimization, chimera creation, and sequence/structural analysis. These steps combine Rosetta- and AI-based methods to improve stability, specificity, and developability, while offering expert support for custom chimeric designs. This integrated approach makes it possible to design highly specific therapeutic antibodies with greater confidence and efficiency.
References
Tae-Sung Oh, et al. Computational design of a single-domain antibody that specifically recognizes WT1 peptide-loaded class I MHC. https://www.biorxiv.org/content/10.1101/2025.07.24.666704v1.full