Antigenicity, Immunogenicity, and Humanness: Concepts and Tools for Therapeutic and Vaccine Design
Introduction
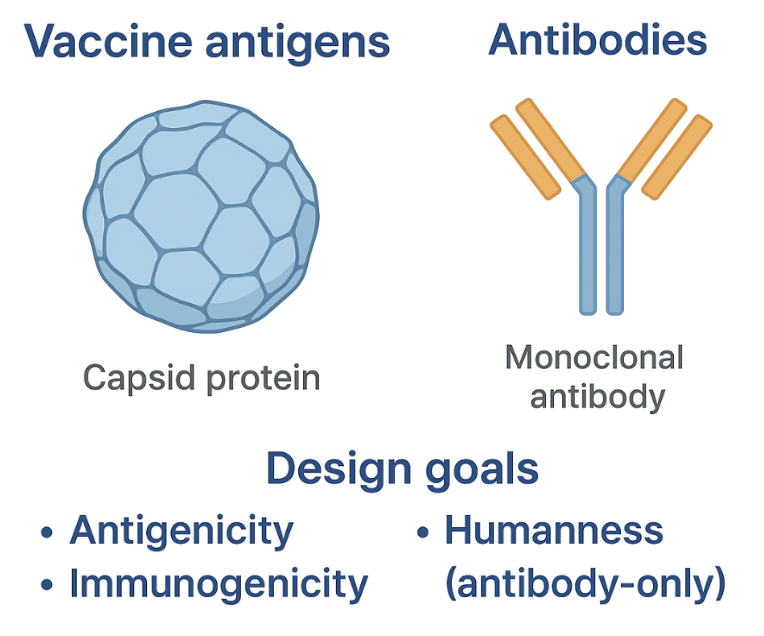
It is important to consider antigenicity and immunogenicity during vaccine design, and additionally, humanness during antibody therapeutic design in order to elicit (in the case of vaccines) or prevent (in the case of therapeutics) an immune response to your designed molecules.
Antigenicity is the ability of a substance to bind to a specific antibody or T cell receptor, while immunogenicity is the ability of a substance to trigger an immune response, including binding to immune receptors and activating immune cells to produce antibodies and a cascade of other immune reactions.
Meanwhile, humanness is an attribute of antibody therapeutics that often describes how similar your design is to other human antibodies, compared to mouse, rabbit, or another species’ antibodies. Therapeutic antibodies are most commonly discovered through animal immunization. However, animal antibody frameworks are often recognized as foreign or antigenic by the human immune system, leading to immunogenicity and specifically an anti-drug antibody (ADA) response. To prevent this, therapeutic antibodies from animal immunization need to undergo a process called humanization. The more effectively humanized an antibody therapeutic—i.e., the greater the humanness of the antibody—the less likely the antibody will be rendered ineffective by the human immune system.
Also, just being more stable can improve immunogenicity. Misfolded or aggregation-prone proteins are often flagged by the immune system, which can complicate both vaccine and therapeutic efficacy. Thus, computational tools must account for sequence, structure, and biophysical properties simultaneously.
Levitate’s Epitope Scan, BioPhi, and Deimmunizer enable researchers to assess vaccine or therapeutic antigenicity, immunogenicity, and humanness, as well as generate design suggestions to humanize or deimmunize an antibody therapeutic. Together, these tools reduce experimental burden by focusing wet-lab assays on the most promising candidates.
Insights
Advances in MHC Binding Prediction for Immunogenicity Assessment
Accurate prediction of peptide–MHC binding is essential for both vaccine and therapeutic design. The NetMHCIIpan-4.0 algorithm represents a major advance, trained on large datasets of naturally eluted ligands (EL). Unlike earlier tools that depended largely on binding affinity assays—which capture only a narrow slice of the antigen presentation process—NetMHCIIpan incorporates large datasets of naturally eluted ligands obtained from immunopeptidomics mass spectrometry. By training on these data, the model learns not only binding motifs but also context-dependent features such as proteolytic antigen processing and peptide length preferences. This shift toward EL-trained models substantially improves the ability to predict which peptides will be presented on the surface of antigen-presenting cells (APCs) and subsequently recognized by T cells, demonstrated by benchmarking on the IEDB. Such accuracy is critical in both therapeutic and vaccine contexts, where immunogenicity hinges on presentation in vivo rather than theoretical binding strength. Levitate’s Epitope Scan applies this machine learning framework and outputs binding score weighted by allele population density, predicted binding affinity in nanomolar IC50, and eluted ligand prediction score.
Deimmunization of Biologics Through Epitope Scanning
Systematically identifying immunogenic epitopes opens broader opportunities to suggest mutations that would reduce immune recognition while preserving protein function. This concept—epitope-driven deimmunization—has become a cornerstone of protein engineering. Levitate’s Deimmunizer builds directly on this paradigm, providing researchers with an automated way to identify MHC-binding peptides in a therapeutic sequence and propose edits that mitigate ADA risk. Deimminizer uses Rosetta to find mutations that are neutral or stabilizing for your protein, but have a lower Epitope Scan score so they have a reduced likelihood of being displayed on APCs. Running the tool is straightforward: users supply a protein structure (PDB) and a residue file defining which positions can be redesigned, while parameters like allele sets, scoring weights, and number of repeats control how broadly the search is explored. The output bundles together both the redesigned protein candidates and supporting files—such as ranked epitope models and residue composition penalties—so researchers can trace how each edit was selected. This not only streamlines deimmunization but also provides transparent, reproducible evidence for every proposed mutation.
Humanness Scoring and Humanization for Antibody Therapeutics
The immunogenicity of therapeutic antibodies is not only a matter of epitope content but also of species origin. The utility of “humanness scores,” computational measures that quantify how similar a given antibody sequence is to the natural human antibody repertoire, with higher humanness correlating strongly with lower ADA incidence in the clinic, has shifted antibody discovery pipelines away from ad hoc grafting methods toward data-driven optimization. Levitate’s BioPhi applies these concepts at scale, allowing users to benchmark their candidates against extensive databases of human antibody sequences and iteratively humanize frameworks without sacrificing stability or binding. BioPhi supports several modes tailored to different design goals: Sapiens for automated humanization using machine learning, CDR grafting for classical framework replacement, Humanness for OASis-based repertoire benchmarking, and Designer for computer-assisted sequence generation. Depending on the mode, the outputs include humanized or designed sequences alongside detailed reports, which provide residue-level annotations and design rationales. These outputs give researchers both actionable sequences and transparent evaluation metrics, enabling them to choose the best balance between humanness, stability, and binding fidelity.
Conclusion
The combined consideration of antigenicity, immunogenicity, and humanness provides a rigorous framework for vaccine or therapeutic design. Advances in computational immunology have transformed these concepts from theoretical concerns into practical, engineerable properties.
By implementing algorithms inspired by decades of academic research, Levitate’s Epitope Scan, BioPhi, and Deimmunizer enable rapid in silico screening, reducing dependence on costly immunoassays and accelerating the path from concept to clinic. For therapeutic antibodies, this means safer and more effective candidates with lower ADA risk. For vaccines, it means higher confidence that designed antigens will elicit robust, population-wide immune responses.