Binder Design
OVERVIEW
Binder Design is a tool for designing protein binders using RoseTTAFold Diffusion (RFDiffusion), developed at the University of Washington. RFDiffusion is a method for protein structure generation (with or without structure specification) useful for a broad range of protein design applications such as vaccines, therapeutics, enzymes, and drug delivery. The RFDiffusion GitHub repository provides many in-depth examples.
The Binder Design tool excels at producing diverse protein structures with highly varied features, possessing the desired structure-based attributes. This diversity enhances the potential for achieving your design goal by expanding beyond limited scaffold libraries from experimental methods.
The resulting structures from Binder Design also exhibit notable advancements in both stability and solubility when refined with ProteinMPNN sequence design and validated with deep learning-based structure prediction, also implemented in Bench.
At Levitate Bio, we not only provide implementation services but also offer valuable insights and guidance to enhance the efficiency of your protein design pipeline.
RUNNING Binder Design
The Binder Design tool in Bench is an intuitive implementation of RFDiffusion for generating de novo binders to an input target structure. Setting up a new job involves:
Selection preparation
-
To begin, open an existing project or create one in Bench. Click on a structure collection in the Structures panel, or load one using the Structure Loader button.
-
Once you have selected the desired structure to run Binder Design on, click the Binder Design button in the Actions panel to open a Binder Design tab.
-
Create a selector for the target protein, domain, or a truncated selection. Recommendations for truncating your target are included in the Choosing a target section of this documentation.
-
Optional: Create a selector the desired binding epitope or site on your target protein, commonly referred to as hotspots. This can be an experimentally determined set of residues or a rationally selected set from structure analysis.
Binder Design job set up
-
Choose the target and optional hotspots selections from the respective dropdown menus.
-
Enter the range of binder lengths to sample. The values are inclusive, so if you select 50 to 100, both 50 residues and 100 residues are possible lengths. To sample one length, enter the same value for the lower and upper bound.
-
Enter the number of designs to generate.
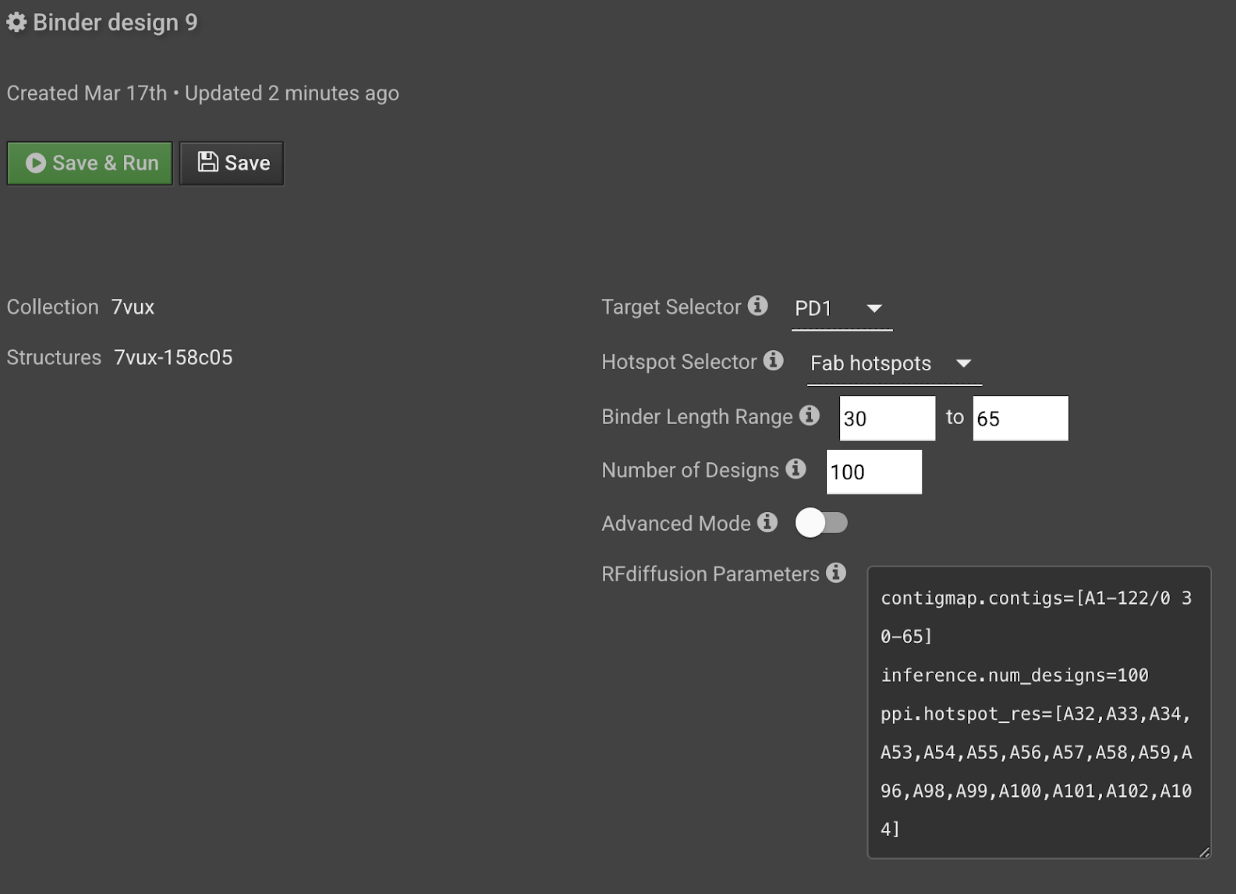
Optional: Review the RFDiffusion parameters automatically created from the above entries. If you have experience with RFDiffusion and wish to edit the RFDiffusion parameters directly without using the above entries, toggle Advanced Mode ON and enter parameters directly in the text field with the following format.
contigmap.contigs=[A1-100/0 50-50]
inference.num_designs=100
ppi.hotspot_res=[A1,A2,A7,A41,A58,A60,A61,A62,A63]
Finally, select Save & Run to start the Binder Design job.
For a comprehensive walk-through, check out the De Novo Binder Design in Bench workflow.
At any step of the pipeline, email support@levitate.bio to receive expert support from our protein design scientists.
References
De novo design of protein structure and function with RFdiffusion